Molecular Signatures
Signature is two-dimensional molecular descriptor based on the molecular graph of a molecule. The vertices of the graph are the set of atoms (or building blocks) in the molecule and the edges are the set of bonds that connect the vertices to one another. A complete list of the signature atom types can be found in the READ-ME file of the translator program.
|
|
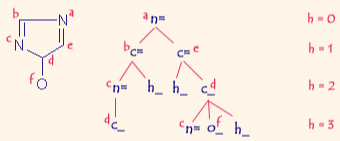
A sample molecular graph of 4H-Imidazol-4-ol rooted on vertex a with 3 levels of branching shown. |
In this manner, a molecule is characterized by a set of unique canonical subgraphs, called signatures, each rooted on a different vertex with a predefined level of branching that describes the local neighborhood up to a distance h away from the root, called the height.
The set of signatures hσ(x) and their occurrence in the molecular graph comprise the molecular descriptors for a molecule. These are expressed as a string of characters corresponding to the canonized subgraph, read in breath-first order. Branch levels are indicated by a set of parenthesis following the parent vertex. (See examples)