Metabolic Engineering using Retrosynthesis
Synthetic biology applied to industrial biotechnology is transforming the way we produce chemicals. However, despite advances in the scale and scope of metabolic engineering, the bioproduction process still remains costly. In order to expand the chemical repertoire for the production of next generation compounds, a major engineering biology effort is required in the development of novel design tools that target chemical diversity through rapid and predictable protocols.
Addressing that goal involves retrosynthesis approaches, which are widely used techniques in chemistry, that explore the chemical biosynthetic space. However, the complexity associated with the large combinatorial retrosynthesis design space has often been recognized as the main challenge hindering the approach.
Our aim is to demonstrate that retrosynthesis can be utilized to engineer chassis
organisms such as E. coli to biosynthesize specific target compounds.
Retrosynthesis algorithms take as input a set of metabolites, for instance the metabolites of a growth medium or the metabolites of a chassis strain model, and the set of target compounds to bioproduce. Ideally the target compounds could be any molecules of the chemical space. The algorithms generate retrosynthesis networks linking the target compound(s) (the source) to the metabolites of the chassis strain (the sink) through reactions. The transformations are applied in the reverse direction from the actual synthesis. In the case of metabolic engineering, retrosynthesis consists of applying reversed biotransformations (i.e., reversed enzymes catalyzed reactions) to a target product, following pathways to substrates that are endogenous to a chassis organism.
To perform such retrosynthesis analysis we developed RetroPath2.0, an automated open source workflow for retrosynthesis a template-based (rule based) method that perform retrosynthesis from chassis to target through an efficient beam-search well-controlled protocol.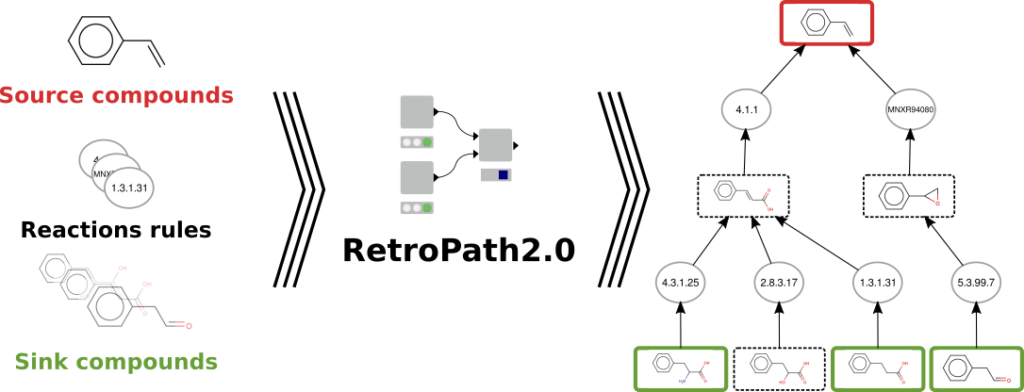
More information about RetroPath2.0 and download link here:
RetroPath2.0: A retrosynthesis workflow for metabolic engineers.
The RetroPath2.0 software is now embeded into the Galaxy toolshed: Galaxy-SynBioCAD.
Retrosynthesis requires to search for enzyme sequences catalyzing reactions. To this end we have developed a series of machine learning methods enabling to screen large databases and predict catalytic activity from sequence.
We have also develop a reinforcement learning retrosynthesis method (RetroPath-RL) based on Monte Carlo Tree Search (MCTS). That method is able to retrieve more and longer pathways than with beam-search.
Details on BioRetroSynth’s background and previous developments (RetroPath v1) can be found here.



