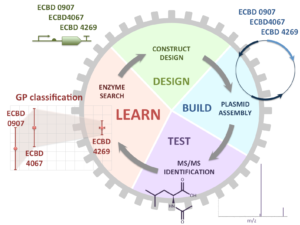
Machine learning for biotechnology applications
Our activities at the interface between machine learning and synthetic biology are twofold. First, we use a variety of learning methods (supervised learning, active learning, and reinforcement learning) to help the synthetic biology process, and secondly, we use synthetic biology to engineer machine learning devices. We have recently started a new ANR-funded project aimed at investigating the cognition capacities of microorganisms. More information can be found on our Artificial Metabolic Networks web page. We are also involved in the newly funded HORIZON BIOS project (cf. press release_final )
We organized a workhop on these topics in June 2024 more information can be found in the following page AI Methods and Models for (Bio)Catalysis and Synthetic Biology Workshop
–
Engineering machine learning devices
- Faulon JL, Ahavi P, Hoang A, Reservoir Computing with Bacteria, bioRxiv 2024 | DOI: 10.1101/2024.09.12.612674
- Faure L, Mollet B, Liebermeister W, Faulon JL. A neural-mechanistic hybrid approach improving the predictive power of genome-scale metabolic models. Nat Commun, 2023, 14, 4669 | DOI: 10.1038/s41467-023-40380-0
- Pandi A, Koch M, Voyvodic PL, Soudier P, Bonnet J, Kushwaha M, Faulon JL. Metabolic Perceptrons for Neural Computing in Biological Systems. Nature Communications, 10: 3880, 2019. | doi: 1038/s41467-019-11889-0
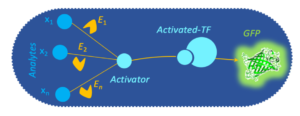
- Voyvodic PL, Pandi A, Koch M, Conejero I, Valjent E, Courtet P, Renard E, Faulon JL*, Bonnet J*. Plug-and-play metabolic transducers expand the chemical detection space of cell-free biosensors. Nature Communications, 10(1):1697, 2019. | doi: 10.1038/s41467-019-09722-9 | PMID: 30979906
–
Reinforcement learning, active learning, and design of experiments
- Pandi A, et al. A versatile active learning workflow for optimization of genetic and metabolic networks. Nat Commun., 2022, 13(1):3876. | DOI: 10.1038/s41467-022-31245-z | PMID: 35790733
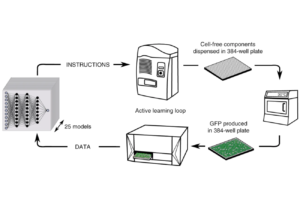
- Borkowski O et al. Large scale active-learning-guided exploration for in vitro protein production optimization. Nature Communications, 11(1): 1872, 2020. | doi: 10.1038/s41467-020-15798-5 | PMID: 32312991
- Koch M, Duigou T, Faulon, JL. Similarity-guided Monte Carlo Tree Search for bio-retrosynthesis. ACS Synthetic Biology, 2019. | doi: 10.1021/acssynbio.9b00447 | PMID: 31841626
- Jervis AJ et al. Machine learning of designed translational control allows predictive pathway optimisation in Escherichia coli. ACS Synthetic Biology, 2018. | doi: 1021/acssynbio.8b00398| PMID: 30563328
- Carbonell P et al. An automated Design-Build-Test-Learn pipeline for enhanced microbial production of fine chemicals. Communications Biology, 1:66, 2018. | doi: 1038/s42003-018-0076-9 | PMID: 30271948
–
Supervised learning for biological sequences
- Carbonell P, Wong J, Swainston N, Takano E, Turner NJ, Scrutton NS, Kell DB, Breitling R, Faulon JL. Selenzyme: Enzyme selection tool for pathway design. Bioinformatics, 2018. | doi: 1093/bioinformatics/bty065
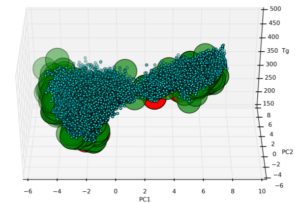
- Mellor J, Grigoras I, Carbonell P, Faulon JL. Semi-supervised Gaussian Process for automated enzyme search. ACS Synthetic Biology, 5(6): 518-528, 2016. | doi: 1021/acssynbio.5b00294
- Carbonell P, Lecointre G, Faulon JL. Origins of specificity and promiscuity in metabolic networks. Journal of Biological Chemistry, 286(51): 43994-44004, 2011. | doi: 1074/jbc.M111.274050
- Misra M, Martin S, Faulon JL*. Graphs: Flexible Representations of Molecular Structures and Biological Networks, in Computational Approaches in Cheminformatics and Bioinformatics, Guha R., Bender, A. Edts, Wiley, 2012. | doi: 1002/9781118131411.ch6
- Carbonell P, Faulon JL. Molecular signatures-based prediction of enzyme promiscuity. Bioinformatics, 26(16): 2012-2019, 2010. | doi: 1093/bioinformatics/btq317
- Faulon JL, Misra M, Martin S, Sale K, Sapra R. Genome scale enzyme-metabolite and drug-target interaction predictions using the signature molecular descriptor. Bioinformatics. 2008 Jan 15;24(2):225-33. Epub 2007 Nov 23. | doi: 1093/bioinformatics/btm580
- Martin S, Brown WM, Faulon JL*. Using product kernels to predict protein interactions. Advances in Biochemical Engineering/Biotechnology, 110:215-245, 2008. | doi: 1007/10_2007_084 | PMID: 17922100
- Brown, W.M., Martin, S., Chabarek, J.P., Strauss, C., Faulon, J.L. Prediction of β-strand packing interactions using the signature product, Journal of Molecular Modeling,12, 355-361, 2006 | doi: 1007/s00894-005-0052-4 | PMID: 16365772
- Martin, S., Roe, D., Faulon, J.L. Predicting protein-protein interactions using signature products, Bioinformatics, 21(2):218-226, 2005. | doi: 1093/bioinformatics/bth483 | PMID: 15319262
–
Supervised learning to predict molecular activity and to design novel molecules.
- Koch M, Duigou T, Carbonell P, Faulon JL. Molecular structures enumeration and virtual screening in the chemical space with RetroPath2.0. Journal of Cheminformatics, 9:64, 2017. | doi: 1186/s13321-017-0252-9
- Planson, A.G., Carbonell, P., Paillard, E., Pollet, N., Faulon, J.L. Compound toxicity screening and structure-activity relationship modeling in Escherichia coli. Biotechnology and Bioengineering, 109(3): 846-850, 2012. | doi: 1002/bit.24356
- Weis DC, Visco DP Jr, Faulon JL*. Data mining PubChem using a support vector machine with the Signature molecular descriptor: classification of factor XIa inhibitors. J Mol Graph Model. 2008 Nov;27(4):466-75. Epub 2008 Aug 27. | doi: 1016/j.jmgm.2008.08.004 | PMID: 18829357
- Brown, W.M., Martin, S., Rintoul, M.D., Faulon, J.L. The Signature Molecular Descriptor. 6. Designing novel polymers with targeted properties using the signature molecular descriptor, Journal of Chemical Information and Modeling, 46(2), 826-835, 2006. | doi: 1021/ci0504521 | PMID: 16563014
- Faulon, J.L., Brown, W.M., Martin, S. Reverse engineering chemical structures from molecular descriptors: how many solutions? Journal of computer-aided molecular design, 19(9-10):637-650, 2005. | doi: 1007/s10822-005-9007-1 | PMID: 16267694
- Faulon, J.L., Visco, D., Churchwell, C.J. The Signature Molecular Descriptor. 2. Enumerating Molcules from their Extended Valence Sequences, Journal of Chemical Information and Computer Sciences, 43 (3), 721 -734, 2003. | doi: 1021/ci020346o | PMID: 12767130
- Faulon, J.L., Visco, D., Pophale, R.S. The Signature Molecular Descriptor. 1. Using Extended Valence Sequences in QSAR and QSPR studies, Journal of Chemical Information and Computer Sciences, 43, 707-720, 2003. | doi: 1021/ci020345w | PMID: 1276712