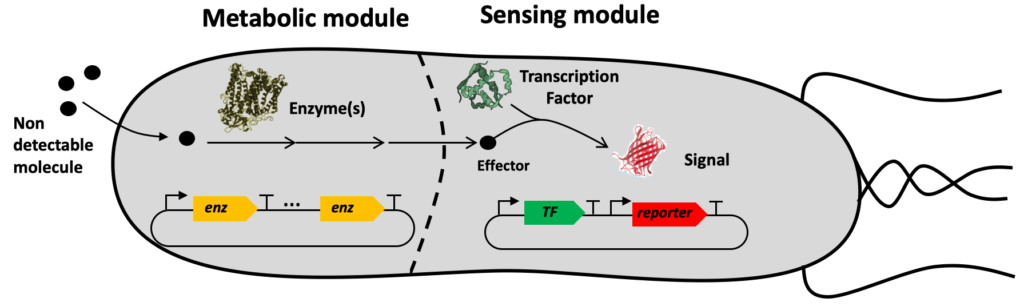
Sensing enabling metabolic pathways (SEMPs) for Biosensors Engineering
Our work on whole-cell and cell-free biosensors is funded by the ANR (French National Funding Agency) through contract ANR-18-CE44-0015. The project named SynBioDiag is a partnership between the CBS (INSERM) and CHU of Montpellier. Our aim is the multiplex detection of prostate cancer biomarkers. Further information on the project can be found at: https://anr.fr/Projet-ANR-18-CE44-0015
We are also funded under the European FET-Open program BioCellPhe. Our contribution is to engineer pathways that upon cancer metastasis biomarker detection trigger the production of chemical signals simultaneously detectable with high sensitivity by surface-enhanced Raman scattering (SERS).
The SynbioDiag and BioCellPhe projects are making use of a technology named ‘Sensing Enabling Metabolic Pathway’ exploiting synthetic networks to expand the portfolio of molecules that are currently known to be detected by natural systems. Our method consists of transforming a priori non-detectable molecules into detectable ones.
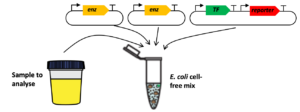
Sensing enabling metabolic pathways can be engineered in whole-cell or in cell-free systems
cell-free
The SEMP technology comprises several computational tools and experimental protocols that can be used together or independently.
- A dataset of detectable molecules (effectors) [8] that activate or repress transcription factors and or riboswitches.
- The RetroPath software [10] to search metabolic pathways transforming any molecule one wishes to detect into an effector.
- The Sensipath web server [12] to visualize SEMPs and get sequence information on transforming enzymes, transcription factors, and riboswitches.
- Protocols to build and test biosensors in whole-cell [2,3,9,13,14] and cell-free systems [1,4,5]
We have benchmarked the technology for a variety of molecules, ranging from detecting central metabolites [14], secondary metabolites [2,9,13], sugars [3,4], chemicals [13] and biomarkers [1,5]. We have showcased the applicability of the SEMP technology in the context of enzyme and metabolic engineering [2,3,4,9,14], environmental pollutant detection [13], and clinical sample analysis [1,5].
REFERENCES
- Soudier P, Ana Zuńiga A, Duigou P, Voyvodic PL, Bazi-Kabbaj K, Kushwaha M, Vendrell JA, Solassol J, Bonnet J, and Faulon JL. PeroxiHUB: A Modular Cell-Free Biosensing Platform Using H2O2 as Signal Integrator. ACS Synthetic Biology, 2022, 11(8):2578-2588 | DOI: 10.1021/acssynbio.2c00138 | PMID: 35913043
- Pandi A, Koch M, Voyvodic PL, Soudier P, Bonnet J, Kushwaha M, Faulon JL. Metabolic Perceptrons for Neural Computing in Biological Systems. Nature Communications, 10: 3880, 2019. | doi: 1038/s41467-019-11889-0
- Castaño-Cerezo S, Fournié M, Urban P, Faulon JL, Truan G. Development of a Biosensor for Detection of Benzoic Acid Derivatives in Saccharomyces cerevisiae. Frontiers in Bioengineering and Biotechnology, 7: 372, 2020. | doi: 10.3389/fbioe.2019.00372 | PMID: 31970152
- Armetta J, Berthome R, Cros A, Pophillat C, Colombo B, Pandi A, Grigoras I. Biosensor-based enzyme engineering approach applied to psicose biosynthesis. Synthetic Biology, 4(1): ysz028, 2019. | doi: 10.1093/synbio/ysz028
- Pandi A, Grigoras I, Borkowski O*, Faulon JL*. Optimizing Cell-Free Biosensors to Monitor Enzymatic Production. ACS Synth Biol. 2019 Aug 16;8(8):1952-1957. | doi: 1021/acssynbio.9b00160 | PMID: 31335131
- Voyvodic PL, Pandi A, Koch M, Conejero I, Valjent E, Courtet P, Renard E, Faulon JL*, Bonnet J*. Plug-and-play metabolic transducers expand the chemical detection space of cell-free biosensors. Nature Communications, 10(1):1697, 2019. | doi: 1038/s41467-019-09722-9| PMID: 30979906
- Koch M, Pandi A, Borkowski O, Cardoso Batista A, Faulon JL*. Custom-made transcriptional biosensors for metabolic engineering. Current Opinion in Biotechnology, 59:78-84, 2019. | doi: 1016/j.copbio.2019.02.016 | PMID: 30921678
- Koch M, Faulon JL*, Borkowski O*. Models for Cell-Free Synthetic Biology: Make Prototyping Easier, Better, and Faster. Frontiers in Bioengineering and Biotechnology, 6: 182, 2018. | doi: 3389/fbioe.2018.00182| PMID: 30555825
- Koch M, Pandi A, Delépine B, Faulon JL*. A dataset of small molecules triggering transcriptional and translational cellular responses. Data in Brief, 17: 1374-1378, 2018. | doi: 1016/j.dib.2018.02.061 | PMID: 29556520
- Trabelsi H, Koch M, Faulon JL*. Building a minimal and generalizable model of transcription-factor based biosensors: showcasing flavonoids. Biotechnology and Bioengineering, 115(9): 2292-2304, 2018. | doi: 1002/bit.26726| PMID: 29733444
- Delépine B, Duigou T, Carbonell P, Faulon JL*. 0: A retrosynthesis workflow for metabolic engineers. Metabolic Engineering, 45: 158-170, 2018. | doi: 10.1016/j.ymben.2017.12.002 | PMID: 29233745
- Libis V, Delépine B, Faulon JL*. Sensing new chemicals with bacterial transcription factors. Current opinion in microbiology, 33: 105-112, 2016. | doi: 1016/j.mib.2016.07.006 | PMID: 27472026
- Delépine B, Libis V, Carbonell P, Faulon JL*. SensiPath: computer-aided design of sensing-enabling metabolic pathways. Nucleic Acids Research, 44: W226-231, 2016. | doi: 1093/nar/gkw305 | PMID: 27106061
- Libis V, Delépine B, Faulon JL*. Expanding biosensing abilities through computer-aided design of metabolic pathways. ACS Synthetic Bioliology, 5(10): 1076-1085, 2016. | doi: 1021/acssynbio.5b00225 | PMID: 27028723
- Fehér T, Libis V, Carbonell P, Faulon JL*. A sense of balance: experimental investigation and modeling of a malonyl-CoA sensor in Escherichia coli. Frontiers in Bioengineering and Biotechnology, 3: 46, 2015. | doi: 3389/fbioe.2015.00046 | PMID: 25905101